Analyzing MD simulations
[1]:
import numpy as np
import matplotlib.pyplot as plt
import chilife as xl
C:\Program Files\Python312\Lib\site-packages\tqdm\auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
[2]:
traj = xl.load_protein('AR1A.pdb', 'AR1A.xtc')
SL = xl.SpinLabel.from_trajectory(traj, 2, spin_atoms=['NN', 'ON'], use_H=True)
print(SL)
Ligand ensemble with 237 members
Name: 2CYR_from_traj
Ligand: CYR
Site: 2
Dihedral definitions:
['N', 'CA', 'CB', 'SG']
['CA', 'CB', 'SG', 'S1L']
['CB', 'SG', 'S1L', 'C1L']
['SG', 'S1L', 'C1L', 'C1R']
['S1L', 'C1L', 'C1R', 'C2R']
[3]:
mosaic = """AABBCC
.DDEE."""
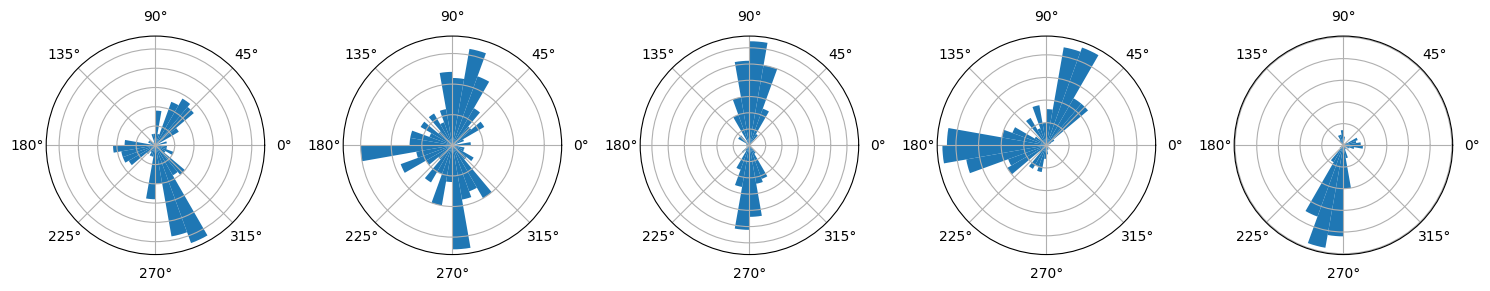
fig, ax_dict = plt.subplot_mosaic(mosaic, figsize=(10, 5), subplot_kw=dict(projection="polar"))
for i, ax in enumerate(ax_dict.values()):
ax.hist(np.deg2rad(SL.dihedrals[:, i]), bins=np.linspace(-np.pi, np.pi, 37))
ax.set_yticklabels([])
xlab = ["0", "45", "90", "135", r"$\rm\pm180$", "-135", "-90", "-45" ]
xlab = [x + r"$\degree$" for x in xlab]
ax.set_xticklabels(xlab)
ax.tick_params(axis='x', which='major', pad=10)
ax.set_title(f"$\\rm\\chi_{i + 1}$")
plt.tight_layout()
plt.show()
[4]:
SL.to_rotlib('CYR', 'Cstom MTSL (R1) rotamer library',
"An MTSL/R1 rotamer library made with well tempered metadynamics")
[5]:
prot = xl.fetch('1ubq')
SL = xl.SpinLabel('CYR', 28, prot)
xl.save(prot, SL)